The construction of the vertebrate brain is dependent on a complex cascade of biological processes that include mitotic division, relocation of migrating neurons, and the extension of dendritic and axonal processes. To gain insight into the molecules that mediate these cellular events my laboratory has relied on unbiased phenotypic screens in mice, the generation of novel transgenic mouse mutants, coupled with the study of human neurodevelopmental disorders. This work has highlighted the importance of the microtubule cytoskeleton and associated proteins in the building of the human brain.
We have shown that mutations in the a-tubulin gene TUBA1A cause lissencephaly a neurodevelopmental disease characterised by a smooth cortex, epilepsy and severe intellectual impairment (Keays et al, Cell 2007). The tubulins are a multi-gene family that encode for the constituents of microtubules; in humans there are 8 α- and 9 β-tubulin genes. Since our discovery that mutations in TUBA1A cause lissencephaly a host of neurological disease states have been associated with the tubulins. For instance, it is now known that variants in TUBB3 cause defects in axon guidance, mutations in TUBA4A cause motor neuron disease, and mutations in the β -tubulin gene TUBB5 cause microcephaly (Breuss et al, Cell Reports 2012). We have built on this work further demonstrating that mutations in the microtubule associated protein EB2 cause the unusual “Michelin tyre” syndrome, and that variants in a microtubule associated kinase (MAST1) cause a rare disorder characterised by an enlarged corpus callosum (Tripathy et al, Neuron 2018). We are interested in understanding how these mutations cause a spectrum of neurodevelopmental diseases and what role these genes play in the developing brain.
Current Research Projects.
Do tubulin genes have specific functions? An emerging theme from our work is that mutations in different tubulin genes give rise to overlapping but distinct disorders. It has been further noted that each tubulin protein has a unique variant C-terminal region and is expressed differentially. These observations support the idea that individual tubulin genes encode functionally distinct polypeptides. Known as the multi-tubulin hypothesis this idea was first proposed more than 40 years ago by Fulton and Simpson, but has not been rigorously testing in a vertebrate system. We are exploiting the power of mouse genetics, coupled with the CRISPR-cas9 genome editing system to test this hypothesis, defining the role of different tubulin genes in the developing brain.
Organoids and tubulin gene mutations. We are interested in understanding how mutations in the tubulin genes cause 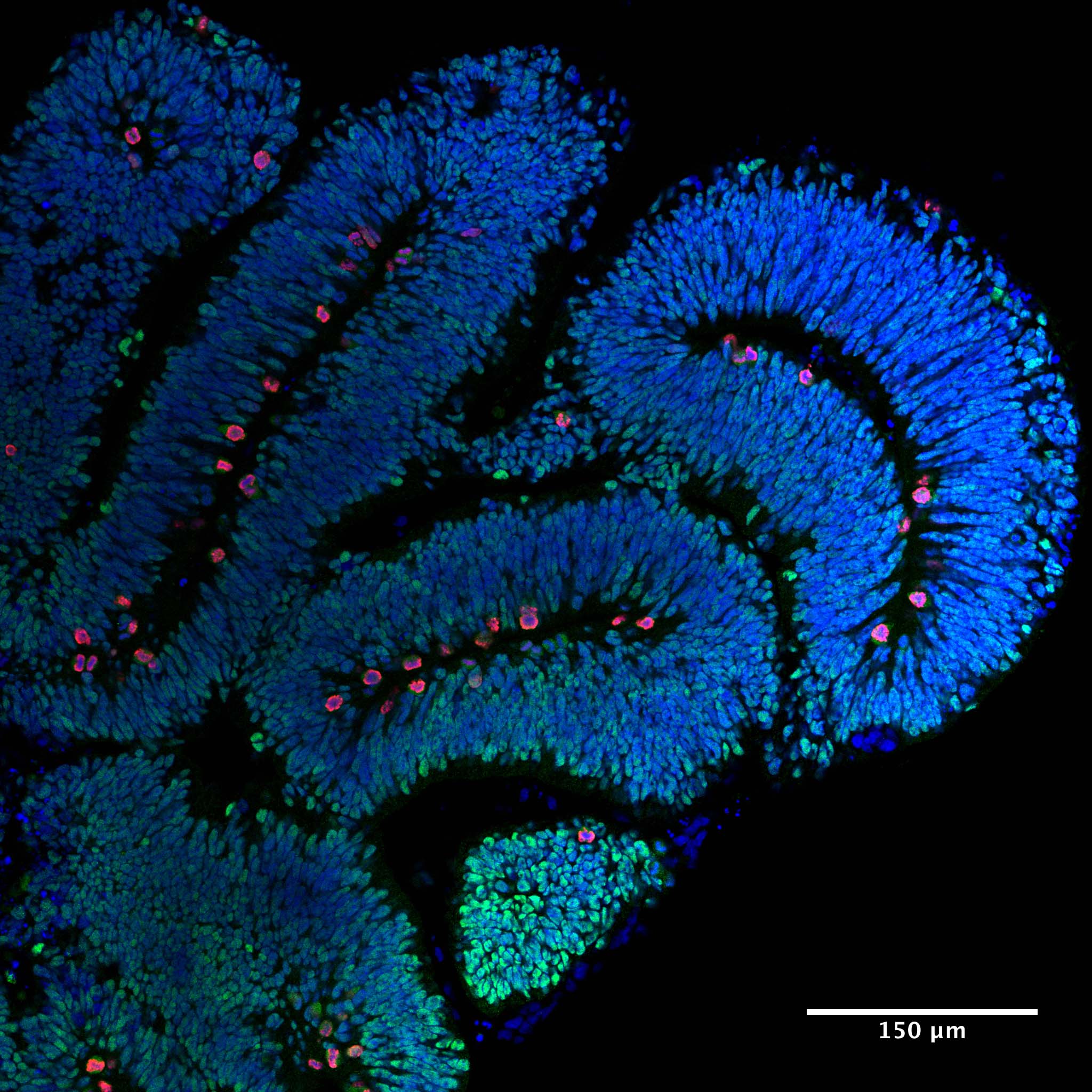 neurodevelopmental disease. To gain mechanistic insight into this question in a human context we are exploiting the power of human cerebral organoids to model the tubulinopathies. Drawing on our established network of clinicians, we generate iPSCs, which are then employed to generate “mini brains” (shown on right). We carefully analyse these organoids alongside controls that have been subject to genetic repair using the CRISP-cas9 genome editing system. We use a variety of methods including brain clearing (iDISCO), light sheet microscopy, live cell imaging, and transcriptomic methods to gain insight into the underlying molecular pathology.
neurodevelopmental disease. To gain mechanistic insight into this question in a human context we are exploiting the power of human cerebral organoids to model the tubulinopathies. Drawing on our established network of clinicians, we generate iPSCs, which are then employed to generate “mini brains” (shown on right). We carefully analyse these organoids alongside controls that have been subject to genetic repair using the CRISP-cas9 genome editing system. We use a variety of methods including brain clearing (iDISCO), light sheet microscopy, live cell imaging, and transcriptomic methods to gain insight into the underlying molecular pathology.
What role does the MAST family play in the developing brain? We have recently shown that microdeletions in MAST1 cause a rare disease characterised by a “mega” corpus callosum, and implicated this gene in autism spectrum disorder. There are four MAST family proteins (MAST1-4), we are largely uncharacterised. Our goal is to define their expression pattern, identify their phosphorylation targets, understand how they regulate microtubule function, and explore the effects of their ablation. To achieve these goals and understand how mutations in the MAST family cause neurological disease we have generated a variety of novel mouse models that recapitulate patient mutations, and reprogrammed iPSCs for organoid development.
Collaborators: Professor Jamel Chelly (Paris), Professor Nick Cowan (NYU), Professor Juergen Knoblich (IMBA).
Funded by: FWF – I2681-B27 – Knock in Mouse Models and Microtubule Associated Cortical Development Disorders, FWF – P29483-B27 – The pathology of the Michelin tire baby Syndrome.